Isolation and Characterization of β-Glucosidase-Producing Bacteria for Compound K Production via Biotransformation
Ⓒ The Korean Environmental Sciences Society. All rights reserved.
This is an Open-Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
Ginsenosides are the major bioactive components of Panax ginseng; however, their limited intestinal absorption has spurred interest in their biotransformation to minor ginsenosides such as compound K. In this study, β-glucosidase–producing bacteria capable of converting major ginsenosides into compound K were isolated and characterized from diverse environmental sources. A total of 28 isolates were screened using esculin hydrolysis and p-nitrophenyl-β-D-glucopyranoside assays, from which strains L66 and K7-1, exhibiting the highest activities, were selected. Based on 16S rRNA gene sequence analysis, L66 and K7-1 were identified as Lactiplantibacillus plantarum and Priestia aryabhattai, respectively. Culture conditions for β-glucosidase production were optimized by evaluating medium type, incubation temperature, and initial pH. Biochemical characterization revealed optimal conditions (45–50°C, pH 7.5) and moderate stability for the crude enzymes. Enzymatic biotransformation experiments conducted under standardized enzyme activity conditions (5.0 U/mL) demonstrated the conversion of protopanaxadiol-type ginsenosides. Notably, as indicated by TLC analysis, strain L66 showed more effective conversion toward compound K and less apparent accumulation of intermediate metabolites than strain K7-1. These findings suggest that L. plantarum L66 is a promising biocatalyst for the eco-friendly, cost-effective production of compound K using crude enzyme systems.
Keywords:
β-Glucosidase, Biotransformation, Compound K, Ginsenosides, Lactiplantibacillus plantarum1. Introduction
Panax ginseng C.A. Meyer has long been recognized as a valuable medicinal plant in East Asia and is widely consumed as a functional food and nutraceutical ingredient. The diverse pharmacological activities of ginseng, including immunomodulatory, anti-inflammatory, antioxidant, and anticancer effects, are primarily attributed to its triterpenoid saponins, known as ginsenosides (Attele et al., 1999; Lü et al., 2009). More than 100 ginsenosides have been identified to date, among which protopanaxadiol (PPD)-type compounds such as Rb1, Rb2, Rc, and Rd are the major constituents of raw and processed ginseng products (Christensen, 2009).
Despite their abundance, most major ginsenosides exhibit limited intestinal absorption due to their high molecular weight and multiple sugar moieties, resulting in low oral bioavailability (Hasegawa et al., 1996; Akao et al., 1998). In the human body, these glycosylated ginsenosides are extensively metabolized by intestinal microbiota into minor deglycosylated metabolites, among which ginsenoside compound K has attracted particular attention. Compound K exhibits superior bioavailability and displays a wide range of biological activities, including anti-inflammatory, antitumor, hepatoprotective, and antidiabetic effects (Yang et al., 2015; Sharma and Lee, 2020). Clinical and pharmacokinetic studies have further demonstrated that compound K represents a major bioactive form detected in plasma following oral administration of ginseng preparations (Kim et al., 2020; Deng et al., 2024).
However, the in vivo formation of compound K via intestinal metabolism is highly variable among individuals, depending on gut microbial composition and enzymatic activity (Akao et al., 1998; Kim et al., 2020). This variability limits the reproducibility and efficacy of ginseng-derived functional products. To overcome this limitation, microbial and enzymatic biotransformation strategies have been extensively investigated as alternative approaches for the controlled production of compound K under defined conditions (Chi and Ji, 2005; Quan et al., 2013; Shin et al., 2015; Upadhyaya et al., 2016).
Among the various biocatalysts, β-glucosidases play a central role in ginsenoside biotransformation by catalyzing the stepwise hydrolysis of β-D-glucosidic linkages attached to the aglycone backbone (Lee et al., 2012; Yan et al., 2016). Efficient conversion of PPD-type ginsenosides into compound K requires coordinated removal of glucose residues at the C-3 and C-20 positions through sequential enzymatic reactions. Accordingly, microorganisms producing highly active and broad-specificity β-glucosidases have been regarded as promising candidates for compound K production (Michlmayr et al., 2010; Zhang et al., 2019).
Previous studies have reported compound K production using purified enzymes, recombinant systems, or genetically engineered strains (Shin et al., 2015; Zhang et al., 2019). Although these approaches enable high conversion efficiency, their practical application is often constrained by high production costs, complex purification processes, and limited operational stability. In contrast, crude enzyme systems derived from natural microorganisms offer operational simplicity, cost-effectiveness, and scalability, making them attractive for industrial bioconversion processes (Garcia et al., 2015; Strahsburger et al., 2017). Nevertheless, systematic screening and characterization of native β-glucosidase-producing strains suitable for crude enzyme-based compound K production remain insufficient.
Lactic acid bacteria and Priestia (formerly Bacillus) species isolated from fermented foods and environmental sources have received increasing attention due to their safety profiles, metabolic versatility, and functional enzyme production capacity (Michlmayr et al., 2010; Paventi et al., 2025). In particular, Lactiplantibacillus spp. (formerly Lactobacillus) has been widely reported to participate in glycoside hydrolysis and functional compound biotransformation during food fermentation, highlighting its potential as a biocatalyst platform (Lee et al., 2012; Paventi et al., 2025).
In this study, β-glucosidase-producing bacteria were isolated from diverse environmental and food-related sources and evaluated for their potential in ginsenoside biotransformation. High-activity strains were identified and characterized, and their biocatalytic performance toward compound K production was systematically investigated. By integrating enzymatic properties with biotransformation behavior, this study aims to establish an environmentally sustainable and economically feasible bioconversion platform for functional ginsenoside production. The findings are expected to contribute to the development of eco-friendly and industrially applicable bioprocesses and to promote the effective utilization of naturally occurring microbial resources in industrial biotechnology.
2. Materials and Methods
2.1. Bacterial strains and culture conditions
In this study, a total of 28 bacterial strains were selected from the laboratory-maintained microbial strain library, including E10 and E33 (isolated from leaf surfaces), G4 (isolated from humus-rich soil), K1, K3, K3-1, K3-4, K4, and K7-1 (isolated from waste feathers), L1, L4, L31, L36, L49, L55, L66, and L77 (isolated from Kimchi), N1, N2, N4, N6, and N7 (isolated from rhizosphere soil), P3 and P15 (isolated from agricultural field soil), W62 and W69 (isolated from dairy products), and Y1 and Y9 (isolated from paddy soil). Unless otherwise stated, strains isolated from Kimchi were cultivated in De Man-Rogosa-Sharpe (MRS) medium (Difco, BD, USA), whereas the remaining strains were cultured in Luria-Bertani (LB) medium (Difco, BD, USA) at 30°C and 200 rpm for 24 h. Short-term storage was performed at 4°C, and long-term preservation was conducted at −80 °C in the presence of 20% (v/v) glycerol.
2.2. Screening and quantitative assay of β-glucosidase activity
Primary qualitative screening for β-glucosidase-producing strains was performed using esculin agar (HiMedia, India) according to the method of Lee et al.(2014). Strains forming dark brown halos were selected as putative β-glucosidase producers.
Quantitative β-glucosidase activity was determined following the method of Matsuura and Obata (1993) with p-nitrophenyl-β-D-glucopyranoside (pNPG) as a chromogenic substrate. Culture broths were centrifuged at 12,000 × g and 4°C for 10 min, and the supernatants were filtered through 0.22 μm syringe filters to obtain crude enzyme extracts. Crude enzyme solution (500 μL) was mixed with 500 μL of 10 mM pNPG prepared in 0.05 M sodium acetate buffer (pH 5.0) and incubated at 40°C for 30 min. The reaction was terminated by adding 1 mL of 0.5 M Na2CO3, and the released p-nitrophenol (pNP) was measured at 405 nm using a UV-Vis spectrophotometer (Ultrospec 4000; Pharmacia Biotech, UK). Heat-inactivated crude enzyme (100°C, 10 min) was used as a negative control. One unit of β-glucosidase activity was defined as the amount of enzyme causing an absorbance change of 0.001 per minute under the assay conditions.
2.3. Molecular identification of selected strains
Strains exhibiting high β-glucosidase activity were identified by 16S rRNA gene sequencing. The 16S rRNA gene was amplified by polymerase chain reaction (PCR) using the universal primers 27F (5’-AGAGTTTGATCCTGGCTCAG-3’) and 1492R (5’-GGTTACCTTGTTACGACTT-3’) (Lane, 1991). The PCR products were purified and sequenced by a commercial sequencing service (Genotech, Daejeon, Republic of Korea). The obtained sequences were analyzed using BLASTN against the NCBI GenBank database to determine their closest phylogenetic relatives. Multiple sequence alignment was performed using Clustal X (version 1.81) (Thompson et al., 1997). Phylogenetic analysis was conducted using MEGA X software, and a phylogenetic tree was constructed based on the neighbor-joining method with 1,000 bootstrap replications to assess branch reliability (Kumar et al., 2018).
2.4. Optimization of β-glucosidase production
To enhance β-glucosidase production, the effects of basal medium type, initial pH, and incubation temperature were systematically evaluated according to the method of Shin et al. (2015). For strain L66, the basal media tested were MRS broth, MRS supplemented with 0.1% (w/v) soluble starch (MRSS), MRS supplemented with 0.1% (w/v) glutinous rice flour (MRSG), yeast extract-based mineral medium (YM), YM supplemented with 0.1% (w/v) soluble starch (YMS), and YM supplemented with 0.1% (w/v) glutinous rice flour (YMG). For strain K7-1, the basal media tested were tryptic soy broth (TSB), TSB supplemented with 0.1% (w/v) soluble starch (TSS), TSB supplemented with 0.1% (w/v) glutinous rice flour (TSG), nutrient broth (NB), NB supplemented with 0.1% (w/v) soluble starch (NBS), NB supplemented with 0.1% (w/v) glutinous rice flour (NBG), Luria-Bertani broth (LB), LB supplemented with 0.1% (w/v) soluble starch (LBS), and LB supplemented with 0.1% (w/v) glutinous rice flour (LBG). The composition of YM medium was yeast extract (1%, w/v), KH2PO4 (0.1%, w/v), MgSO4 (0.05%, w/v), KCl (0.05%, w/v), FeSO4 (0.01%, w/v), and MnCl2 (0.05%, w/v). All media were adjusted to pH 7.0 prior to sterilization.
Following medium screening, the effects of initial pH (4.0-9.5) and incubation temperature (25-35°C) on β-glucosidase production were further investigated under otherwise identical cultivation conditions.
2.5. Biochemical characterization of crude β-glucosidase
Crude enzyme preparations were used for biochemical characterization. Optimal temperature was determined over the range of 25-60°C at 5°C intervals. Optimal pH was evaluated using 0.05 M buffers ranging from pH 3.0 to 11.0, including sodium acetate/sodium citrate (pH 3.0-5.5), potassium phosphate (pH 6.0-8.0), Tris-HCl (pH 7.0-9.0), and glycine-NaOH/bicarbonate-NaOH (pH 9.0-11.0). Thermal stability was assessed by incubating enzymes at 4-100°C for 1 h prior to activity measurement. pH stability was determined after incubating enzyme samples in buffers of different pH values at 30°C for 1 h, followed by residual activity measurement.
2.6. Enzymatic bioconversion of ginsenosides
Red ginseng concentrate (Korea Ginseng Corp., Korea) was used as the substrate. The concentrate was diluted with sterile distilled water to obtain a total ginsenoside concentration of 5 mg/mL and filtered through a 0.22 μm syringe filter. Crude enzyme and substrate solutions were mixed at a ratio of 1:1 (v/v) and incubated under the optimal pH and temperature determined in Section 2.5 at 100 rpm for 1-7 days in the dark. To ensure comparability among reactions, the crude enzyme was added to achieve a final β-glucosidase activity of 5.0 U/mL in the reaction mixture, as determined by the pNPG assay described in Section 2.2. Samples were collected at regular intervals for product analysis.
2.7. TLC analysis of biotransformation products
Biotransformation products were analyzed using silica gel 60 F254 TLC plates (Merck, Germany) according to the method of Chi et al. (2005). Reaction mixtures (5 mL) were extracted with saturated n-butanol (2 mL) by vigorous mixing, and the upper organic phase containing ginsenosides was collected for TLC analysis. The developing solvent system consisted of chloroform: methanol: water (65:35:10, v/v/v), and the lower (chloroform-rich) phase was used for development after phase separation. Plates were sprayed with 10% sulfuric acid in ethanol and heated at 110°C for 5 min. Compound K was identified by comparison with authentic standards (AceEMzyme, Korea). For semi-quantitative comparison of band intensity, TLC plates were photographed under identical illumination and camera settings, and densitometric analysis was performed using ImageJ (NIH, USA). Band intensities were quantified as integrated optical density after local background subtraction and were normalized to the total lane signal to enable relative comparison among reaction times and between strains.
2.8. Data analysis
All experiments were performed in triplicate. Data are presented as mean ± standard deviation (SD). Statistical significance was evaluated using one-way analysis of variance followed by Tukey’s post hoc test (p < 0.05).
3. Results and Discussion
3.1. Screening of β-glucosidase-producing bacteria
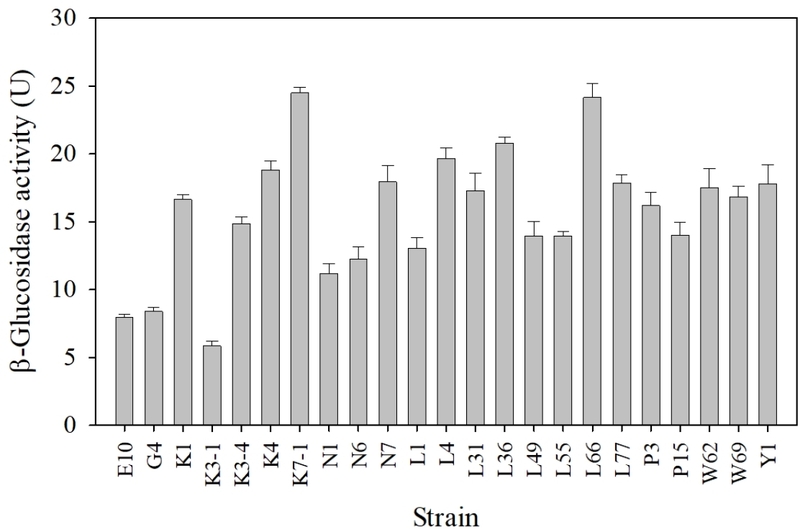
A total of 28 isolates were screened for β-glucosidase activity using an esculin hydrolysis assay. On esculin agar, β-glucosidase-positive strains produce characteristic dark brown to black zones resulting from the formation of iron-esculetin complexes following esculin cleavage, a reaction that has been widely applied as a preliminary screening method for β-glucosidase producers (Lee et al., 2014). Except for strains E33, K3, N2, N4, and Y9, all tested isolates formed distinct dark brown zones surrounding the colonies on esculin agar, indicating positive β-glucosidase activity in the primary screening assay (Fig. 1). The esculin-positive isolates were subsequently subjected to quantitative β-glucosidase activity measurement using pNPG as a chromogenic substrate. Hydrolysis of pNPG releases p-nitrophenol, which can be quantified sensitively after alkalinization, enabling reliable comparison among strains (Strahsburger et al., 2017; Kim et al., 2018). Among the tested isolates, strains L66 and K7-1 exhibited the highest β-glucosidase activities and were therefore selected for further investigation (Fig. 2). The combined use of plate-based screening and quantitative assays enhanced selection reliability by minimizing false positives associated with colony pigmentation, diffusion variability, and growth-related artifacts (Strahsburger et al., 2017).
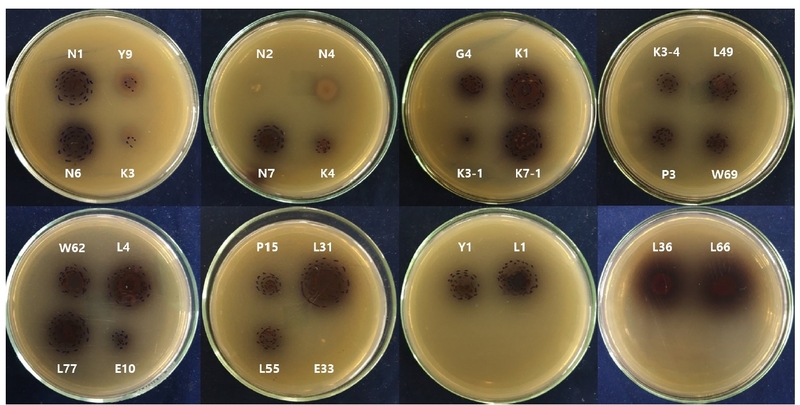
Primary qualitative screening of β-glucosidase-producing strains on esculin agar plates. Dark brown to black halos surrounding colonies indicate positive β-glucosidase activity.
3.2. Molecular identification and phylogenetic analysis
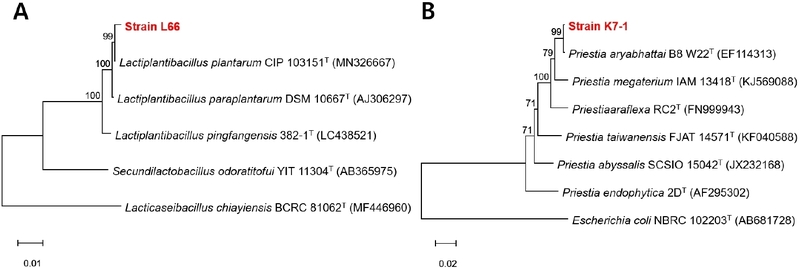
The taxonomic identities of strains L66 and K7-1 were determined based on 16S rRNA gene sequence analysis. Pairwise nucleotide alignment revealed that strain L66 shared 99% sequence identity with Lactiplantibacillus plantarum, while strain K7-1 exhibited 99% sequence identity with Priestia aryabhattai. These results indicate a high degree of nucleotide conservation between the selected isolates and their closest reference strains. Phylogenetic analysis of the 16S rRNA gene sequences further supported the sequence similarity results. As shown in Fig. 3, strain L66 clustered with the type strain of L. plantarum, whereas strain K7-1 grouped closely with the type strain of P. aryabhattai. Because 16S rRNA gene sequencing has limited discriminatory power among closely related species within the L. plantarum complex, strain L66 was conservatively identified at the species level as L. plantarum. Both isolates were clearly separated from closely related reference taxa, confirming their distinct phylogenetic positions.
3.3. Optimization of culture conditions for β-glucosidase production
To enhance enzyme productivity, the effects of basal medium type, incubation temperature, and initial pH were systematically evaluated for strains L66 and K7-1. Carbon source composition and nitrogen availability are critical determinants of β-glucosidase expression, as polysaccharides and oligosaccharides frequently act as metabolic inducers (Garcia et al., 2015; Neesa et al., 2020).
For strain L66, MRSG supported the highest enzyme production (Fig. 4A), whereas for strain K7-1, maximal activity was observed in NBS medium (Fig. 4B). These results indicate strain-specific substrate utilization patterns and regulatory responses, consistent with previous reports on heterogeneous induction mechanisms among microbial β-glucosidases (Garcia et al., 2015). Similar induction effects of starch-based substrates have been reported in solid-state and submerged fermentations of β-glucosidase-producing microorganisms (Neesa et al., 2020).
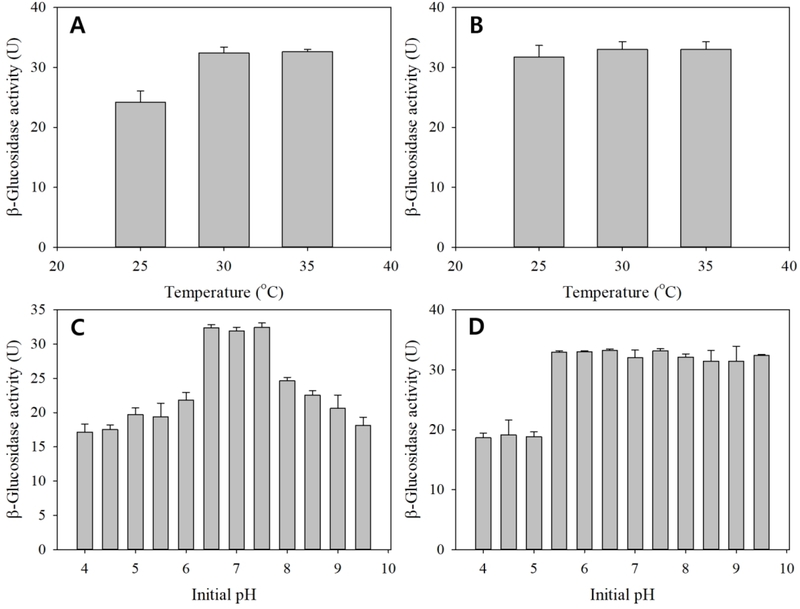
Incubation temperature and initial pH also significantly influenced enzyme production (Fig. 5). Both strains exhibited maximal production at 30-35°C (Fig. 5A and B), which is consistent with optimal enzyme secretion observed in mesophilic β-glucosidase producers (Yan et al., 2016; Neesa et al., 2020). With respect to initial pH, strain L66 showed an optimal range under mildly acidic to neutral conditions (Fig. 5C), whereas K7-1 maintained relatively high productivity across a broader pH range (Fig. 5D). Such strain-dependent pH responses have been attributed to differences in regulatory networks, membrane transport systems, and extracellular enzyme stability (Michlmayr et al., 2010; Li et al., 2025).
3.4. Physicochemical characterization of crude β-glucosidase
The biochemical properties of crude β-glucosidase preparations were evaluated by determining temperature and pH optima, as well as thermal and pH stabilities. Both enzymes exhibited maximal catalytic activity at approximately 45-50°C (Fig. 6A and B), which is consistent with reported optima of microbial β-glucosidases used for glycoside hydrolysis and biotransformation processes (Lee et al., 2012; Yan et al., 2016; Li et al., 2025). The high β- glucosidase activity and favorable catalytic properties observed for strain L66 are in agreement with previous reports highlighting the important role of L. plantarum in food fermentation and functional compound biotransformation (Paventi et al., 2025). Notably, enzyme activity from strain K7-1 declined sharply at elevated temperatures, whereas L66-derived enzyme retained higher activity over a wider temperature range, suggesting superior structural stability of the L66 enzyme. Similar strain-dependent thermal tolerance has been reported for crude β-glucosidase preparations (Michlmayr et al., 2010; Garcia et al., 2015). Both enzymes showed highest activities near neutral to slightly alkaline pH (pH 7.5; Fig. 6C and D), consistent with buffer-dependent modulation of β-glucosidase activity through ionization changes in catalytic residues and substrate-binding microenvironments (Yan et al., 2016; Strahsburger et al., 2017). Stability assays further distinguished the two preparations: the L66 enzyme retained substantial residual activity after moderate thermal pretreatment, whereas the K7-1 enzyme exhibited a narrower thermal stability range before rapid inactivation (Fig. 7A and B). Both enzymes maintained appreciable activity over a broad pH range, although stability decreased under strongly alkaline conditions (Fig. 7C and D), which is relevant for prolonged bioconversion processes in which pH drift can occur (Garcia et al., 2015; Yan et al., 2016). Comparable thermostable and pH-tolerant β-glucosidases with industrial potential have been reported in metagenomic and environmental studies, supporting the practical relevance of such physicochemical properties (Jeilu et al., 2024).
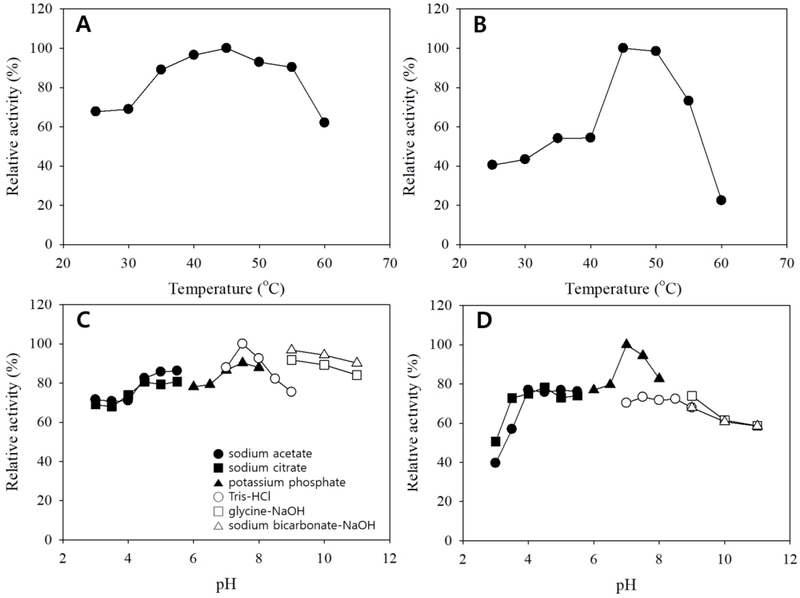
Effects of reaction temperature and pH on β-glucosidase activity from strains L66 and K7-1. A, L66 (temperature); B, K7-1 (temperature); C, L66 (pH); D, K7-1 (pH).
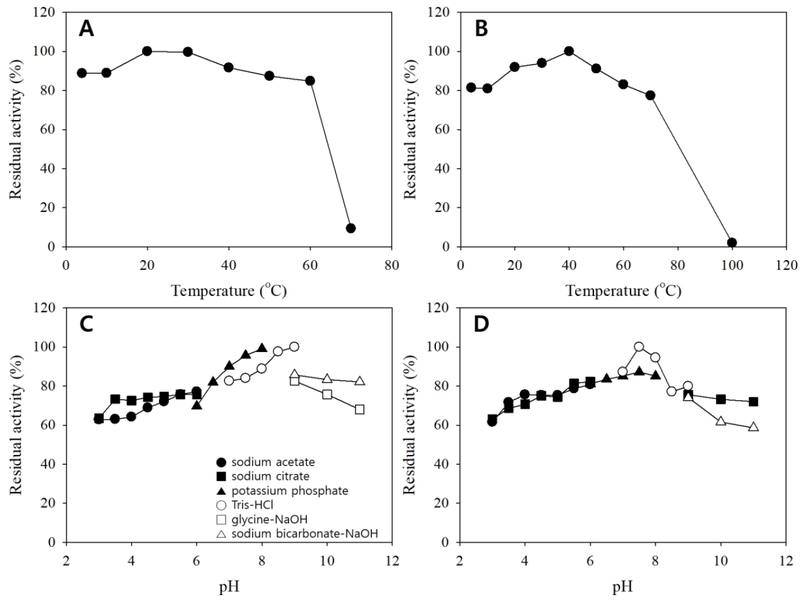
Thermal and pH stabilities of β-glucosidase from strains L66 and K7-1. A, L66 (thermal stability); B, K7-1 (thermal stability); C, L66 (pH stability); D, K7-1 (pH stability).
These enzymatic characteristics are directly relevant to ginsenoside biotransformation because compound K is produced via sequential deglycosylation of major PPD-type ginsenosides (Lee et al., 2012; Shin et al., 2015), and enzymes with broad operational stability can enhance conversion efficiency in crude enzyme systems (Zhang et al., 2019; Sharma and Lee, 2020). Accordingly, the crude β-glucosidase preparations from strains L66 and K7-1 represent practical biocatalyst candidates for compound K production with operational simplicity compared with purified or recombinant enzyme systems.
3.5. TLC analysis of ginsenoside biotransformation
Time-dependent changes in ginsenoside profiles during biotransformation by strains L66 and K7-1 were analyzed using TLC (Fig. 8). In both strains, gradual reduction of major PPD-type ginsenosides (Rb1, Rb2, and Rc) was accompanied by a progressive increase in the compound K band, indicating sequential deglycosylation.
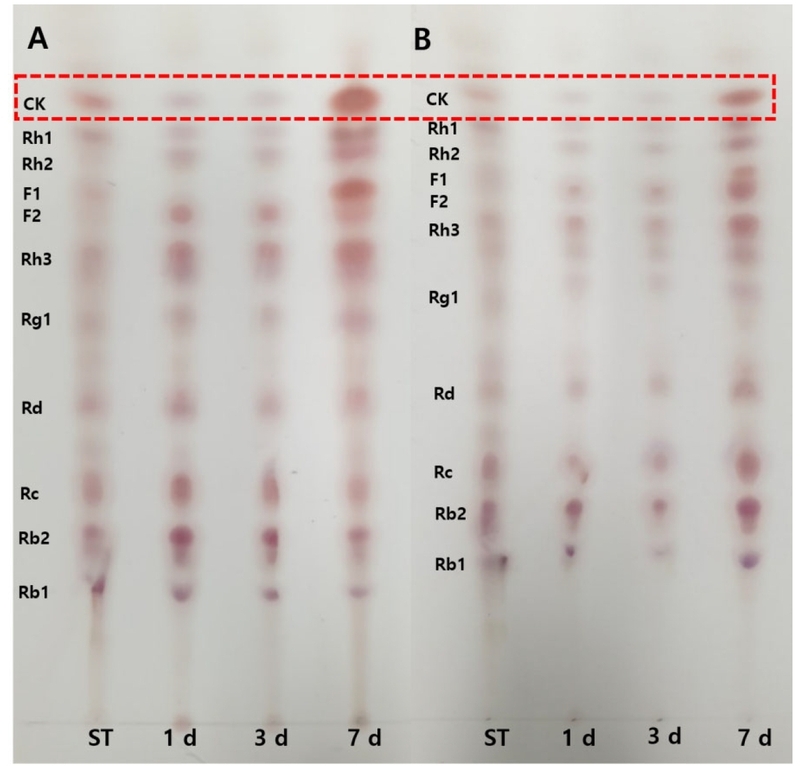
Time-course TLC profiles of ginsenoside biotransformation by strains L66 (A) and K7-1 (B). Compound K was identified by comparison with authentic standards. ST, authentic standards; CK, compound K.
In strain L66 (Fig. 8A), the compound K band intensified markedly by day 7, whereas bands corresponding to precursor ginsenosides, particularly Rc and Rb2, were substantially weakened, and intermediate metabolites such as Rd and F2 decreased at later time points. This pattern suggests that strain L66 efficiently catalyzed both intermediate and terminal hydrolysis steps leading to compound K accumulation. In contrast, strain K7-1 (Fig. 8B) also exhibited time-dependent compound K formation; however, relatively stronger residual bands corresponding to Rc, Rb2, and Rd were observed even after 7 days. This persistence indicates that, although K7-1 possesses β-glucosidase activity toward major ginsenosides, its conversion efficiency for late-stage steps appears lower than that of L66 under the tested conditions.
The observed biotransformation patterns are consistent with the well-established sequential deglycosylation pathway of PPD-type ginsenosides, in which Rb1, Rb2, and Rc are converted to Rd and F2, followed by the formation of compound K (Chi and Ji, 2005; Chi et al., 2005; Quan et al., 2013; Upadhyaya et al., 2016). Similar TLC profiles, characterized by progressive disappearance of precursor bands and accumulation of compound K, have been reported for efficient compound K-producing food-derived and intestinal microorganisms (Quan et al., 2013; Upadhyaya et al., 2016). Notably, the progressive reduction of intermediate metabolites in strain L66 suggests the presence of a highly coordinated β-glucosidase system capable of catalyzing both intermediate and terminal hydrolysis reactions. Such a “complete-type” deglycosylation system has been associated with high compound K productivity in previous studies using Lactobacillus, Bifidobacterium, and Paecilomyces strains (Yang et al., 2015; Kim et al., 2020).
From a physiological perspective, compound K is known to be generated predominantly through intestinal microbial metabolism and is rarely detected in the absence of bacterial hydrolysis (Akao et al., 1998; Deng et al., 2024). The TLC pattern observed for strain L66 closely resembles that of highly active intestinal bacteria, in which limited accumulation of intermediates and preferential compound K formation are observed. Therefore, strain L66 may function as an effective in vitro model for intestinal-type ginsenoside metabolism.
Collectively, these results demonstrate clear strain-dependent differences in ginsenoside biotransformation efficiency, with strain L66 exhibiting more efficient precursor conversion and dominant compound K accumulation than strain K7-1.
4. Conclusions
In this study, β-glucosidase-producing bacteria capable of converting major ginsenosides into compound K were systematically screened from diverse environmental sources. Among 28 isolates, strains L66 and K7-1 exhibited the highest β-glucosidase activities and were identified as L. plantarum and P. aryabhattai, respectively, based on 16S rRNA gene sequence analysis. Optimization of culture conditions revealed that β-glucosidase production by both strains was strongly influenced by medium type, incubation temperature, and initial pH, with strain-specific response patterns. Biochemical characterization demonstrated that the crude enzymes from both strains exhibited favorable temperature and pH optima, as well as moderate thermal and pH stabilities, supporting their applicability in bioconversion processes. Enzymatic biotransformation experiments using standardized enzyme activities (5.0 U/mL) confirmed that both strains were capable of converting PPD-type ginsenosides into compound K. Consistent with the biotransformation analysis, strain L66 demonstrated more efficient conversion of intermediate metabolites and preferential accumulation of compound K compared with strain K7-1, indicating a more coordinated deglycosylation system. Overall, this study demonstrates that L. plantarum L66 represents a promising biocatalyst for the efficient production of compound K using crude enzyme preparations, offering operational simplicity and practical applicability compared with purified or recombinant enzyme systems. The findings provide a practical basis for the potential development of environmentally friendly and cost-effective bioprocesses for functional ginsenoside production. Further studies focusing on process scale-up, enzyme stabilization, and reaction engineering will contribute to the industrial application of this bioconversion system.
Acknowledgments
This work was supported by a 2-Year Research Grant of Pusan National University.
REFERENCES
-
Akao, T., Kida, H., Kanaoka, M., Hattori, M., Kobashi, K., 1998, Intestinal bacterial hydrolysis is required for the appearance of compound K in rat plasma after oral administration of ginsenoside Rb1 from Panax ginseng, J. Pharm. Pharmacol., 50(10), 1155-1160.
[https://doi.org/10.1111/j.2042-7158.1998.tb03327.x]

-
Attele, A. S., Wu, J. A., Yuan, C. S., 1999, Ginseng pharmacology: Multiple constituents and multiple actions, Biochem. Pharmacol., 58(11), 1685-1693.
[https://doi.org/10.1016/S0006-2952(99)00212-9]

-
Chi, H., Ji, G. E., 2005, Transformation of ginsenosides Rb1 and Re from Panax ginseng by food microorganisms, Biotechnol. Lett., 27(11), 765-771.
[https://doi.org/10.1007/s10529-005-5632-y]

-
Chi, H., Kim, D. H., Ji, G. E., 2005, Transformation of ginsenosides Rb2 and Rc from Panax ginseng by food microorganisms, Biol. Pharm. Bull., 28(11), 2102-2105.
[https://doi.org/10.1248/bpb.28.2102]

-
Christensen, L. P., 2009, Ginsenosides chemistry, biosynthesis, analysis, and potential health effects, Adv. Food Nutr. Res., 55, 1-99.
[https://doi.org/10.1016/S1043-4526(08)00401-4]

-
Deng, M. S., Huang, S. T. Z., Xu, Y. N., Shao, L., Wang, Z. G., Chen, L. J., Huang, W. H., 2024, In vivo pharmacokinetics of ginsenoside compound K mediated by gut microbiota, PLoS One, 19(8), e0307286.
[https://doi.org/10.1371/journal.pone.0307286]

-
Garcia, N. F. L., Santos, F. R. S., Gonçalves, F. A., Paz, M. F., Fonseca, G. G., Leite, R. S. R., 2015, Production of β-glucosidase on solid-state fermentation by Lichtheimia ramosa in agroindustrial residues: Characterization and catalytic properties of the enzymatic extract, Electron. J. Biotechnol., 18(4), 314-319.
[https://doi.org/10.1016/j.ejbt.2015.05.007]

-
Hasegawa, H., Sung, J. H., Matsumiya, S., Uchiyama, M., 1996, Main ginseng saponin metabolites formed by intestinal bacteria, Planta Med., 62(5), 453-457.
[https://doi.org/10.1055/s-2006-957938]

-
Jeilu, O., Alexandersson, E., Johansson, E., Simachew, A., Gessesse, A., 2024, A Novel GH3-β-glucosidase from soda lake metagenomic libraries with desirable properties for biomass degradation, Sci. Rep., 14, 10012.
[https://doi.org/10.1038/s41598-024-60645-y]

- Kim, J. H., Lee, S. Y., Park, H. J., Choi, Y. J., 2018, Fermentative transformation of ginsenosides by a combination of probiotic Lactobacillus helveticus and Pediococcus pentosaceus, Kor. J. Microbiol., 54(4), 436-441.
-
Kim, J. K., Choi, M. S., Jeung, W., Ra, J., Yoo, H. H., Kim, D. H., 2020, Effects of gut microbiota on the pharmacokinetics of protopanaxadiol ginsenosides Rd, Rg3, F2, and compound K in healthy volunteers treated orally with red ginseng, J. Ginseng Res., 44(4), 611-618.
[https://doi.org/10.1016/j.jgr.2019.05.012]

-
Kumar, S., Stecher, G., Li, M., Knyaz, C., Tamura, K., 2018, MEGA X: Molecular evolutionary genetics analysis across computing platforms, Mol. Biol. Evol., 35(6), 1547-1549.
[https://doi.org/10.1093/molbev/msy096]

- Lane, D. J., 1991, 16S/23S rRNA sequencing, in: Stackebrandt, E., Goodfellow, M. (eds.), Nucleic acid techniques in bacterial systematics, John Wiley and Sons, New York, 115-175.
-
Lee, K. W., Han, N. S., Kim, J. H., 2012, Purification and characterization of β-glucosidase from Weissella cibaria 37, J. Microbiol. Biotechnol., 22(12), 1705-1713.
[https://doi.org/10.4014/jmb.1206.06007]

-
Lee, S., Lee, Y. H., Park, J. M., Bai, D. H., Jang, J. K., Park, Y. S., 2014, Bioconversion of ginsenosides from red ginseng extract using Candida allociferrii JNO301 isolated from Meju, Mycobiology, 42(4), 368-375.
[https://doi.org/10.5941/MYCO.2014.42.4.368]

-
Li, L., Liu, H., Liu, T., Mi, J., Cai, R., Xu, H., 2025, β-Glucosidase: Progress from basic mechanism to frontier application, Fermentation, 11(10), 588.
[https://doi.org/10.3390/fermentation11100588]

-
Lü, J. M., Yao, Q., Chen, C., 2009, Ginseng compounds: An Update on their molecular mechanisms and medical applications, Curr. Vasc. Pharmacol., 7(3), 293-302.
[https://doi.org/10.2174/157016109788340767]

-
Matsuura, M., Obata, A., 1993, β-Glucosidases from soybeans hydrolyze daidzin and genistin, J. Food Sci., 58(1), 144-147.
[https://doi.org/10.1111/j.1365-2621.1993.tb03231.x]

-
Michlmayr, H., Schümann, C., da Silva, N. M., Kulbe, K. D., del Hierro, A. M., 2010, Isolation and basic characterization of a β-glucosidase from a strain of Lactobacillus brevis isolated from a malolactic starter culture, J. Appl. Microbiol., 108(2), 550-559.
[https://doi.org/10.1111/j.1365-2672.2009.04461.x]

- Neesa, L., Islam, R., Jahan, N., Zohora, U. S., Rahman, M. S., 2020, Optimization of culture conditions and reaction parameters of β-glucosidase from a new isolate of Bacillus subtilis (B1), J. Appl. Biotechnol. Rep., 7(3), 152-158.
-
Paventi, G., Di Martino, C., Coppola, F., Iorizzo, M., 2025, β-Glucosidase activity of Lactiplantibacillus plantarum: a key player in food fermentation and human health, Foods, 14(9), 1451.
[https://doi.org/10.3390/foods14091451]

-
Quan, L. H., Kim, Y. J., Li, G. H., Choi, K. T., Yang, D. C., 2013, Microbial transformation of ginsenoside Rb1 to compound K by Lactobacillus paralimentarius, World J. Microbiol. Biotechnol., 29(6), 1001-1007.
[https://doi.org/10.1007/s11274-013-1260-1]

-
Sharma, A., Lee, H. J., 2020, Ginsenoside compound K: Insights into recent studies on pharmacokinetics and health-promoting activities, Biomolecules, 10(7), 1028.
[https://doi.org/10.3390/biom10071028]

-
Shin, K. C., Choi, H. Y., Seo, M. J., Oh, D. K., 2015, Compound K production from red ginseng extract by β-glycosidase from Sulfolobus solfataricus supplemented with α-L-arabinofuranosidase from Caldicellulosiruptor saccharolyticus, PLoS One, 10(12), e0145876.
[https://doi.org/10.1371/journal.pone.0145876]

-
Strahsburger, E., Lopez de Lacey, A. M., Marotti, I., Di Gioia, D., Biavati, B., Dinelli, G., 2017, In vivo assay to identify bacteria with β-glucosidase activity, Electron. J. Biotechnol., 30, 83-87.
[https://doi.org/10.1016/j.ejbt.2017.08.010]

-
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., Higgins, D. G., 1997, The CLUSTAL X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools, Nucleic Acids Res., 25(24), 4876-4882.
[https://doi.org/10.1093/nar/25.24.4876]

-
Upadhyaya, J., Kim, M. J., Kim, Y. H., Ko, S. Y., Park, H. W., Kim, M. K., 2016, Enzymatic formation of compound-K from ginsenoside Rb1 by enzyme preparation from cultured mycelia of Armillaria mellea, J. Ginseng Res., 40(2), 105-112.
[https://doi.org/10.1016/j.jgr.2015.05.007]

-
Yan, F. Y., Xia, W., Zhang, X. X., Chen, S., Nie, X. Z., Qian, L. C., 2016, Characterization of β-glucosidase from Aspergillus terreus and its application in the hydrolysis of soybean isoflavones, J. Zhejiang Univ. Sci. B, 17(6), 455-464.
[https://doi.org/10.1631/jzus.B1500317]

-
Yang, X. D., Yang, Y. Y., Ouyang, D. S., Yang, G. P., 2015, A Review of biotransformation and pharmacology of ginsenoside compound K, Fitoterapia, 100, 208-220.
[https://doi.org/10.1016/j.fitote.2014.11.019]

-
Zhang, R., Huang, X. M., Yan, H. J., Liu, X. Y., Zhou, Q., Luo, Z. Y., Tan, X. N., Zhang, B. L., 2019, Highly selective production of compound K from ginsenoside Rd by hydrolyzing glucose at C-3 glycoside using β-glucosidase of Bifidobacterium breve ATCC 15700, J. Microbiol. Biotechnol., 29(3), 410-418.
[https://doi.org/10.4014/jmb.1808.08059]

Department of Life Science and Environmental Biochemistry, Pusan National University bboobbi@pusan.ac.kr
Department of Life Science and Environmental Biochemistry, Pusan National University alswn940809@pusan.ac.kr
Department of Life Science and Environmental Biochemistry, Pusan National University jtm0620a@pusan.ac.kr
Department of Life Science and Environmental Biochemistry, Pusan National University wjdwnsgh9470@pusan.ac.kr
∙ Researcher. O-Mi LeeAvian Disease Division, Animal and Plant Quarantine Agency lomi78@korea.kr
Department of Life Science and Environmental Biochemistry, Pusan National University shjoo@pusan.ac.kr